Set environment
Code
suppressMessages (suppressWarnings (source ("../run_config_project_sing.R" )))show_env ()
You are working on Singularity: singularity_proj_encode_fcc
BASE DIRECTORY (FD_BASE): /data/reddylab/Kuei
REPO DIRECTORY (FD_REPO): /data/reddylab/Kuei/repo
WORK DIRECTORY (FD_WORK): /data/reddylab/Kuei/work
DATA DIRECTORY (FD_DATA): /data/reddylab/Kuei/data
You are working with ENCODE FCC
PATH OF PROJECT (FD_PRJ): /data/reddylab/Kuei/repo/Proj_ENCODE_FCC
PROJECT RESULTS (FD_RES): /data/reddylab/Kuei/repo/Proj_ENCODE_FCC/results
PROJECT SCRIPTS (FD_EXE): /data/reddylab/Kuei/repo/Proj_ENCODE_FCC/scripts
PROJECT DATA (FD_DAT): /data/reddylab/Kuei/repo/Proj_ENCODE_FCC/data
PROJECT NOTE (FD_NBK): /data/reddylab/Kuei/repo/Proj_ENCODE_FCC/notebooks
PROJECT DOCS (FD_DOC): /data/reddylab/Kuei/repo/Proj_ENCODE_FCC/docs
PROJECT LOG (FD_LOG): /data/reddylab/Kuei/repo/Proj_ENCODE_FCC/log
PROJECT REF (FD_REF): /data/reddylab/Kuei/repo/Proj_ENCODE_FCC/references
Import data
Code
= file.path ("region_nuc" , "fcc_astarr_macs" ,"summary" = dir (txt_fdiry)for (txt in vec){cat (txt, " \n " )}
K562.hg38.ASTARR.macs.KS91.input.rep_all.max_overlaps.q5.tsv
K562.hg38.ASTARR.macs.KS91.input.rep_all.union.q5.tsv
Code
### set file directory = file.path (FD_RES, "region_nuc" , "fcc_astarr_macs" , "summary" )= "*tsv" = file.path (txt_fdiry, txt_fname)### get files = Sys.glob (txt_fglob)= basename (vec_txt_fpath)### read files = lapply (vec_txt_fpath, function (txt_fpath){= read_tsv (txt_fpath, show_col_types = FALSE )return (dat)names (lst) = vec_txt_fname### assign data = lst
Check data
Code
= lst_dat_region_import= lst[[1 ]]print (dim (dat))fun_display_table (head (dat))
chr1
10038
10405
chr1:10038-10405
0.523161
367
chr1
14282
14614
chr1:14282-14614
0.578313
332
chr1
16025
16338
chr1:16025-16338
0.587859
313
chr1
17288
17689
chr1:17288-17689
0.625935
401
chr1
28934
29499
chr1:28934-29499
0.771681
565
chr1
115429
115969
chr1:115429-115969
0.381481
540
Code
= lst_dat_region_import= lst[[2 ]]print (dim (dat))fun_display_table (head (dat))
chr1
10015
10442
chr1:10015-10442
0.522248
427
chr1
14253
14645
chr1:14253-14645
0.573980
392
chr1
16015
16477
chr1:16015-16477
0.541126
462
chr1
17237
17772
chr1:17237-17772
0.614953
535
chr1
28903
29613
chr1:28903-29613
0.759155
710
chr1
30803
31072
chr1:30803-31072
0.501859
269
Arrange table
Helper function to map label
Code
= function (vec_txt_input){= c ("overlap" , "union" )= c ("ATAC (Overlap)" , "ATAC (Union)" )= fun_str_map_detect (return (vec_txt_output)### test function fun_str_map_label ("ASTARR_overlap" )
'ATAC (Overlap)'
Combine tables
Code
### Concatenate tables = lst_dat_region_import= bind_rows (lst, .id = "Label" )### Update labels = dat %>% dplyr:: mutate (Label = fun_str_map_label (Label))### assign and show = datprint (dim (dat))print (table (dat$ Label))fun_display_table (head (dat, 3 ))
[1] 396894 7
ATAC (Overlap) ATAC (Union)
150042 246852
ATAC (Overlap)
chr1
10038
10405
chr1:10038-10405
0.523161
367
ATAC (Overlap)
chr1
14282
14614
chr1:14282-14614
0.578313
332
ATAC (Overlap)
chr1
16025
16338
chr1:16025-16338
0.587859
313
Explore data
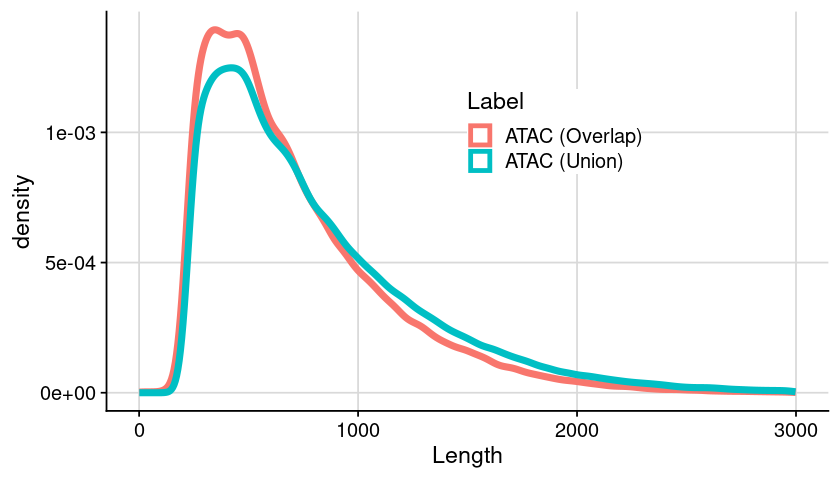
Distribution of region length (double check)
Code
### arrange table = dat_region_arrange= dat %>% :: mutate (Length = ChromEnd - ChromStart) %>% :: filter (Length < 3000 )### generate plot = ggplot (dat, aes (x = Length, color = Label)) + geom_density (linewidth= 2 ) + theme_cowplot () + background_grid () + theme (legend.position = "inside" , legend.position.inside = c (0.5 , 0.7 ),legend.background = element_rect (fill = "white" )### show plot options (repr.plot.height= 4 , repr.plot.width= 7 )print (gpt)
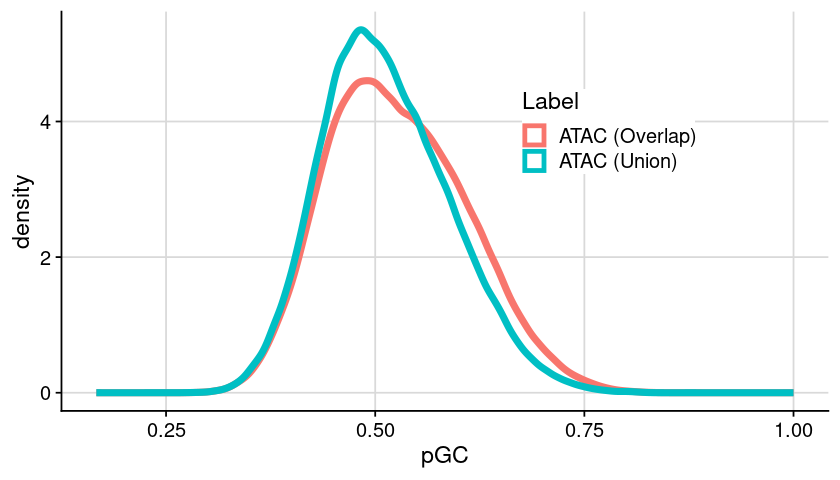
Distribution of GC content
Code
### init = dat_region_arrange### generate plot = ggplot (dat, aes (x = pGC, color = Label)) + geom_density (linewidth= 2 ) + theme_cowplot () + background_grid () + theme (legend.position = "inside" , legend.position.inside = c (0.6 , 0.7 ),legend.background = element_rect (fill = "white" )### show plot options (repr.plot.height= 4 , repr.plot.width= 7 )print (gpt)
Export results
Summarize information
Code
= function (x, ...) {= density (x, ...)= obj$ x[which.max (obj$ y)]return (num)
Code
### compute counts and info = dat_region_arrange= dat %>% group_by (Label) %>% summarise (Mean = mean (pGC), Mode = fun_mode_continuous (pGC),.groups= "drop" )= dat### build a named vector of new legend labels, each with two lines: ### original label ### Count = 12345 = dat_region_annot= dat %>% :: mutate (line1 = Label,line2 = paste ("Mean =" , round (Mean, 2 ), " " ),line3 = paste ("Mode =" , round (Mode, 2 ), " " ),combined = paste0 (line1, " \n " , line2, " \n " , line3)%>% set_names (.$ combined, .$ Label) }= vecprint (vec_txt_region_label)
ATAC (Overlap)
"ATAC (Overlap)\nMean = 0.53 \nMode = 0.49 "
ATAC (Union)
"ATAC (Union)\nMean = 0.51 \nMode = 0.48 "
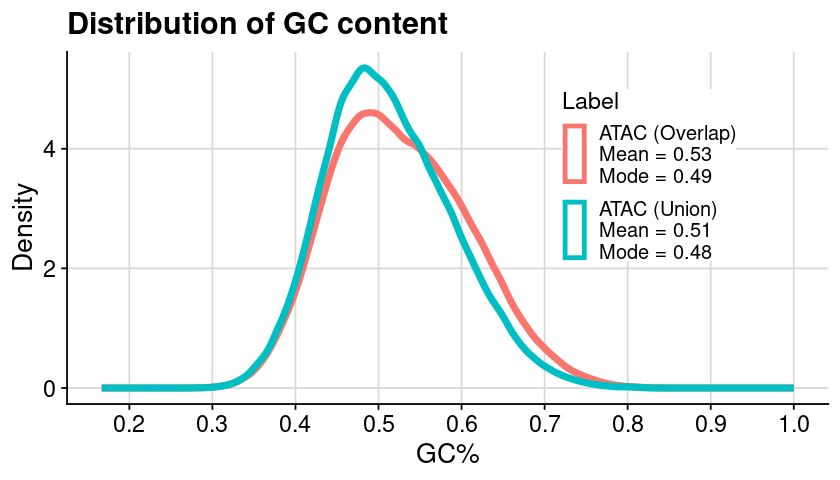
Create plot
Code
### set text size <- theme (title = element_text (size = 16 ),axis.title = element_text (size = 16 ),axis.text = element_text (size = 14 ),legend.title = element_text (size = 14 ),legend.text = element_text (size = 12 )
Code
### init = dat_region_arrange### generate plot = ggplot (dat, aes (x = pGC, color = Label)) + geom_density (linewidth= 2 ) + theme_cowplot () + background_grid () + labs (x = "GC%" , y = "Density" , title = "Distribution of GC content" ) + scale_x_continuous (breaks = seq (0 , 1 , by = 0.1 )) + scale_y_continuous (breaks = seq (0 , 5 , by = 2 )) + scale_color_discrete (labels = vec_txt_region_label) + + theme (legend.position = "inside" , legend.position.inside = c (0.65 , 0.65 ),legend.background = element_rect (fill = "white" ),legend.key.spacing.y = unit (0.3 , 'cm' )### assign plot = gpt### show plot options (repr.plot.height= 4 , repr.plot.width= 7 )print (gpt)
Save plot
Code
= "./" = "fig.region.fcc_astarr_macs_input.distribution.gc_content.png" = file.path (txt_fdiry, txt_fname)ggsave (txt_fpath, gpt_export, height = 4 , width = 7 , units = "in" )= "./" = "fig.region.fcc_astarr_macs_input.distribution.gc_content.svg" = file.path (txt_fdiry, txt_fname)ggsave (txt_fpath, gpt_export, height = 4 , width = 7 , units = "in" )